What's next?
Sun, Feb 8, 2026
Read in 10 minutes
After years of work, a moment of post-PhD clarity to assess the research landscape and a chance to pivot in a new direction. But to what?
Why did I do a PhD?
PhDs attract bright, motivated young people. I should know, I used to be one. That’s not to say I’m particularly bright, but I certainly used to be young. These people are attracted for many reasons: altruism; curiosity; ambition; maybe naïveté, but I can only speak for myself.
I was working in industry in a job that I liked, with good co-workers and good prospects. Why give that up? Aside from the fact that I wanted to live somewhere new, and a PhD gives medium-term license to move somewhere fun (like Austin), I wanted to become an expert at something. My job at Apple had introduced me to the fast-paced world of corporate R&D. We helped develop materials across Apple’s entire product line, from metalloceramic coatings on cosmetic surfaces to battery materials and even precious metal alloys like the limited edition gold Hermès Apple watch. I really only had two complaints:
- Big companies can afford to hire anyone. If I wanted to, say, learn to use a complicated instrument like a TEM, why would they go to the length of training me? They can afford to just hire an actual expert. That leads to my second point.
- Almost everyone had advanced degrees of their own. I don’t want to contribute to a culture of credentialism, but it can be hard to make your voice heard when you’re the least qualified person in the room.
Even for people with technical experience, it’s nearly impossible to know the reality of a particular field until you’re enmeshed in the research. From the outside, it’s easy to underestimate the challenges, and maybe even overestimate the usefulness or generality of most research. The reality is most research is incredibly focused and often targeted towards a very small community of researchers. I say this not to be cynical, contrarian, or obnoxious.
What did I learn?
For me, a PhD was absolutely worthwhile, and sure, I achieved my goal of ‘gaining expertise’. But the value of the degree has far more to do with how I learned than what I learned. Almost everything was self-taught. This began slowly, if ambitiously. Like many young students I dove headfirst into dense piles of literature, and quickly became bogged down by language that the authors assumed their readers were fluent in. Over time, that fluency came, and after half a year or so of reading, I could finally skim the materials science and solid state physics literature without being constantly baffled by the terms before me.
I learned not to be intimidated by dense hyperspecific language, long pages of equations, or complicated codebases. When you don’t have the time and energy to digest these results they can seem incomprehensible, but given enough time and layers of abstraction, ordinary people can build incredible things. A dissertation doesn’t get written in a day, and neither does a large codebase.
Second, I learned that many scientific fields are more similar than they first appear. Indeed, sometimes it feels that about half of the experimental sciences is little more than fixing vacuum leaks, tracing wires, and knowing the correct order to restart computers. Even data analysis is surprisingly similar across disciplines. Spectroscopy looks awfully similar whether you’re using a STEM to measure X-Ray spectra of single atoms or whether you’re using a telescope to measure infrared specta from faraway star. In both cases, the challenge lies with the fact that the signal is incredibly weak. Across domains the same problems and methods appear over and over again.
What do I want? To be an academic? To work in industry?
More than anything, I want to work on the cutting edge of technology. Whether this happens in industry or academia shouldn’t be important. Students are often told that the day-to-day reality of corporate R&D is uninspiring. According to this trope, academics practice a more ‘pure’ and ‘intellectually rigorous’ version of a field. All their counterparts in industry do, we’re told, is throw labor and money until the problems fix themselves.
This may be too generous to academics, some of whom are salesmen just as skilled as any that can be found in industry. Still others have at least one fantastic hammer, with which everything else looks like a nail. In my opinion, too much academic research involves buying a sophisticated tool and training people to press ‘go’. This arrangement outsources some of the most important innovation to equipment vendors.
What options are left? Open, foundational corporate research scarcely exists. There is no modern equivalent to the Bell Labs of yesteryear. In my opinion, two options remain.
- ‘Deep’ tech companies making highly technical innovations. Of course, without the ability to scrutinize their claims, we have little choice but to take them at their word. I don’t want to sound overly critical here. Startups are innately vulnerable, often needing to defend against entrenched, well-resourced competitors. I can think of several startups in the hardware space that have made very impressive advancements. It’s barely worth mentioning that I can think of many more that over-promised and under-delivered. These companies must promise enough to attract investment, but remain vague enough to not give away important details about their approach, a balancing act that can easily descend into hype or deception.
- National labs, where much of the open hardware and instrumentation development occurs. This is where I know that there is groundbreaking work going on, not necessarily because of how groundbreaking it is, but because I know about it.
Okay, so I would like to work at a ‘deep’ tech company or a National lab. But what do I want to pivot into?
What new research areas?
What do I want in a R&D job? The following attributes come to mind. I want:
- Technical work. The less hand-waving the better.
- Hands-on-ness. If data is being acquired, I want to have some part of it. If data science is being done, I’d like to contribute there, too. If instrumentation is being developed, I’d like to contribute however I can.
- Impact. It would be nice to solve a problem that benefits people, rather than hurts them. Let’s get specific about a few new research directions.
Cryo Electron Microscopy
In terms of impact, it’s hard to think of a research area more valuable than the life sciences. The fields upstream of biological electron microscopy - infectious pathology, drug development, neuroscience, etc. - have a much more direct effect upon our well-being than the promise of better LEDs or marginally faster microprocessors. Even better, the CryoEM methods behind this research involve really cool technology - single particle analysis, electron diffraction, tomography, direct electron detectors - that I have experience with and can make meaningful contributions to.
There is, however, a catch. I recently had a conversation with a ML researcher friend who works at a major player in the emerging field of function-driven protein design. I told him that, although I’m interested in moving towards structural biology, I’m afraid that AlphaFold has made routine structure determination something of a solved problem, at least for proteins. He agreed with my concerns, and explained that his group invests considerably more money into computational structural prediction than they do into characterization experiments. I want to be careful here. Just because (some) protein structures can be accurately predicted and because at least one domain expert considers structure determination a ‘solved problem’ doesn’t mean that cryoEM is suddenly irrelevant.
Predictive models excel at generating plausible static folds, but they remain limited in describing conformational ensembles, heterogeneous assemblies, transient complexes, and more. CryoEM, in my opinion, will be essential for investigating those regimes that are poorly captured by sequence-based predictors. I expect ML models to serve as priors for increasingly complicated systems, ultimately shifting the role of cryoEM within the field. If routine, single-protein structure prediction becomes less central, the experimental frontier will move towards increasingly sophisticated questions. The bottleneck will no longer be with technical proficiency in data collection and reconstruction, but with formulating testable hypotheses and in the design of experiments. For someone like myself with minimal formal training in biology, that shift will likely make it more difficult to break in to the field.
This says nothing of the rapidly-growing application of cryoEM techniques to the inorganic sciences. A recent paper of mine, which is currently under review, applies single particle analysis (SPA) to study the structure of octahedral rhenium cluster complexes (although my study was actually carried out at elevated, rather than cryogenic, temperatures). I argue that SPA, which has long been used to study biological structures, is also applicable to inorganic materials, provided that they have sufficient structural stability and homogeneity.
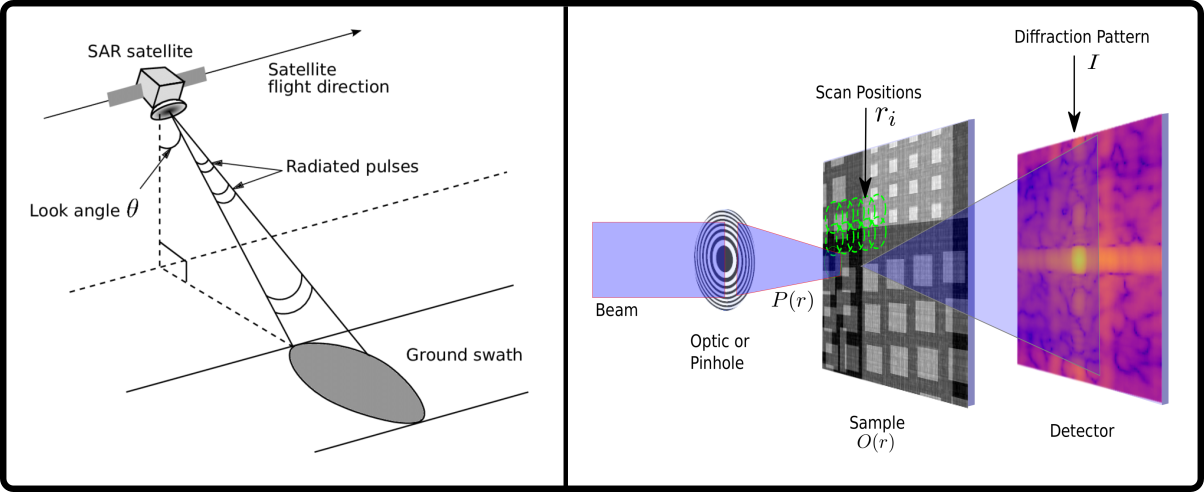
The exchange goes both ways. Techniques developed for materials science are beginning to have an impact in structural biology, too. Scanning TEM methods have been used for single particle analysis in biology. Ptychography, which was developed for inorganic electron microscopy but came into its own on synchrotron beamlines, holds promise for certain cryogenic applications in both materials science and biology. Single-crystal electron diffraction, once considered a method best practiced by an aging class of old-school microscopists, has been reborn as the hot new field of microED.
Looking forward, there is another advancement that I expect to transform biological and matsci electron microscopy alike: automation.
High-Throughput Synthesis and Characterization
Materials synthesis has, historically, been an incredibly tedious endeavor. A single student or postdoc, working for years, is lucky to develop a reliable synthesis method for a new material. Robotic laboratories have the potential to upend and accelerate synthesis in the physical sciences. The idea has been around for a long time, but the devil’s in the details. At a high level there are three requirements:
- A synthesis method to produce large numbers of high-quality samples covering your parameter space.
- A characterization method capable of accurately profiling the desired properties.
- Data science methods to predict properties and generate new candidate samples.
The laboratory engineering and instrumentation hurdles are associated with 1) and 2). This is particularly true of electron microscopy, where sample preparation is notoriously difficult. TEM operation is also no small matter. Other characterization methods that have been employed in a high-throughput context (XRD, XPS, SAXS) have been much simpler to automate.
There is no ‘off the shelf’ solution for this type of work. Advancing high-throughput EM will require significant work in instrumentation, sample handling, robotics, controls, and data science. Some fields, like the neuroscience sub-field of connectomics, have seen impressive applications of high-throughput tomography. These advances shouldn’t occur in isolation. Software developed to control and automate biological microscopes should be flexible and extensible enough to be applied to STEMs. The field is sure to grow, and I’m excited about where it will lead us.
Other fields
Earlier, I mentioned that disparate fields of the physical sciences employ methods that are surprisingly similar. Training in one begets familiarity, or at least conceptual comfort, with the other. Even though I’m no plasma physicist, I’m drawn by the impressive development of detector tech, in-situ instrumentation, and algorithmic processing that goes on at particle beamlines, including at XFELs. Optical microscopy, too, has been transformed by the emergence of methods such as Fourier ptychography.
Concluding thoughts
The amount of fantastic research going on all over the world is humbling, even daunting. I’m highly driven to make a contribution wherever I can. One’s career can be long and winding. Professor Warner didn’t even touch an electron microscope, the tool that would ultimately define his early career, until his second post-doc. More important than proficiency with any specific method is determination to learn more and write, write, write. So, to revisit the question from the beginning of this post: what am I looking for? I wish to do really exciting R&D in industry (preferably a so-called ‘deep tech’ startup), or work on methods and instrumentation at a National lab. To anyone who has read this far, wish me luck.